Introduction
In this article, we will solve LeetCode 187: Repeated DNA Sequences using Java. This is a popular string problem that tests your understanding of the sliding window technique and efficient use of hash-based data structures.
DNA sequences are composed of four characters:
- A (Adenine)
- C (Cytosine)
- G (Guanine)
- T (Thymine)
The goal is to identify all 10-letter-long substrings that appear more than once in a given DNA string.
You can try solving the problem directly on LeetCode here:
https://leetcode.com/problems/repeated-dna-sequences/
Problem Statement
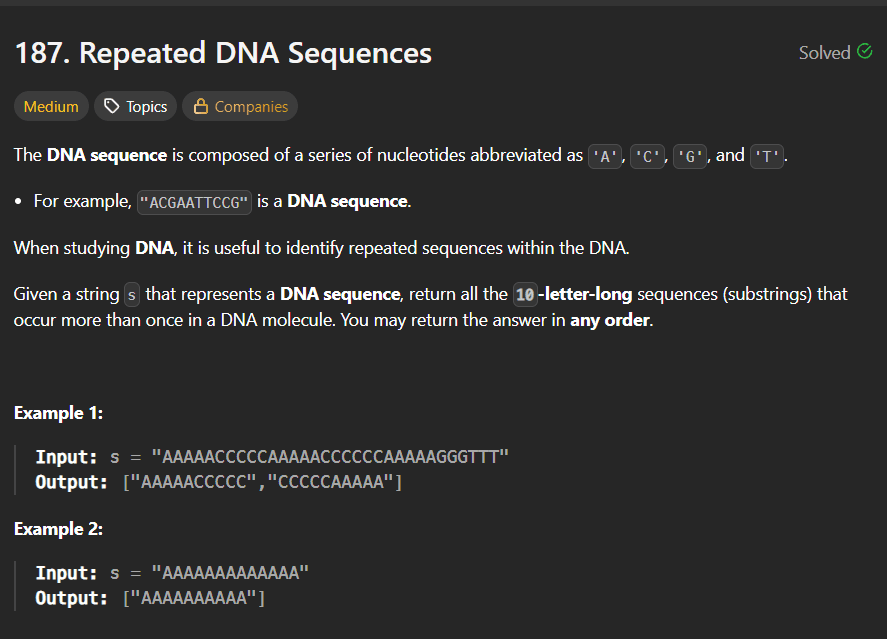
Given a string s that represents a DNA sequence, return all the 10-letter-long substrings that occur more than once.
Example 1
Input:
s = "AAAAACCCCCAAAAACCCCCCAAAAAGGGTTT"
Output:
["AAAAACCCCC", "CCCCCAAAAA"]
Example 2
Input:
s = "AAAAAAAAAAAAA"
Output:
["AAAAAAAAAA"]
Key Observations
- We only need substrings of fixed length 10.
- The maximum length of the string can be up to 10^5.
- A brute-force solution checking all substrings multiple times would be inefficient.
- This problem can be solved efficiently using a sliding window and hash-based data structures.
Approach 1: Sliding Window with HashSet (Given Solution)
Idea
- Use two pointers (
iandj) to maintain a sliding window. - Build a substring of size 10 dynamically.
- Store previously seen substrings in a HashSet.
- If a substring is already present in the set:
- Check if it is already in the result list.
- If not, add it to the result list.
- Slide the window forward and continue.
Java Code (Your Implementation)
Explanation
- The variable
tesmaintains the current substring. msstores all previously seen substrings of length 10.- If a substring already exists in
ms, we manually check whether it has already been added to the result list. - This avoids duplicate entries in the final output.
Time Complexity
- Sliding through the string: O(n)
- Checking duplicates in the result list: O(n) in the worst case
- Overall worst-case complexity: O(n²)
Space Complexity
- HashSet storage: O(n)
Limitation of Approach 1
The manual duplicate check using a loop inside the result list introduces unnecessary overhead. This makes the solution less efficient.
We can improve this by using another HashSet to automatically handle duplicates.
Approach 2: Optimized Solution Using Two HashSets
Idea
- Use one HashSet called
seento track all substrings of length 10. - Use another HashSet called
repeatedto store substrings that appear more than once. - Iterate from index 0 to
s.length() - 10. - Extract substring of length 10.
- If adding to
seenfails, it means it has appeared before. - Add it directly to
repeated.
This removes the need for a nested loop.
Optimized Java Code
Why This Approach is Better
- No manual duplicate checking.
- Cleaner and more readable code.
- Uses HashSet properties efficiently.
- Each substring is processed only once.
Time Complexity (Optimized)
- Single traversal of the string: O(n)
- Substring extraction of fixed length 10: O(1)
- Overall time complexity: O(n)
Space Complexity
- Two HashSets storing substrings: O(n)
Conclusion
LeetCode 187 is a classic example of combining the sliding window technique with hash-based data structures.
- The first approach works but has unnecessary overhead due to manual duplicate checks.
- The second approach is more optimal, cleaner, and recommended for interviews.
- Always leverage the properties of HashSet to avoid redundant checks.
This problem highlights the importance of choosing the right data structure to optimize performance.